
Aamir Khan
Ph.D. Bioinformatics
2
Publications
4 yr 5 mo
Duration
Research Thesis
Title
Study on computational based approach for genome-wide prediction of AMP and molecular markers in buffalo
Objectives
1. To develop artificial intelligence-based algorithm for AMP prediction using transcriptomic and genomic data. 2. To discover polymorphic molecular markers in buffalo. 3. To develop microRNA gene atlas of buffalo. 4. To develop web genomic resource for buffalo.
Abstract
Water buffalo (Bubalus bubalis), belonging to Bovidae family is economically important animal. It is one of the most important farm animals, lives in hot and humid regions where climatic conditions are favorable for prevalence of infectious diseases. But interestingly, there is a less occurrence of diseases along with lesser deleterious effect of diseases on buffalo which indicates their stronger innate immunity to fight against infection. Buffaloes are found to express many AMPs such as defensins, cathelicidins, and hepcidin, which play an important role in neutralizing the invading pathogens. This study provides a faster, improved and species-specific AMP/non-AMP candidate prediction server, BufAMPpred (http://login1.cabgrid.res.in:5030/) based on an ensemble of CNN+LSTM deep learning neural network architecture with improved prediction accuracy (98.72% accuracy, 99.79% sensitively and 98.68% specificity) in comparison to existing tools/servers till date. This server also facilitates analyses up to 500 sequences at a time and also linked with BufAMPdb (http://backlin.cabgrid.res.in/buffampdb/) which is the collection of all candidate AMPs (61711) and non-AMPs (2529971) predicted for buffalo along with their corresponding gene information. We also report the first comprehensive, holistic and user-friendly web genomic resource of buffalo (BuffGR) accessible at http://backlin.cabgrid.res.in/buffgr/, that catalogues 6028881 SNPs and 613403 InDels extracted from the set of 31 buffalo tissues. We found a total of 3727122 SNPs and 634124 InDels distributed in the four breeds of buffalo (Murrah, Bangladesh, Jaffarabadi and Egyptian) with reference to Mediterranean breed. It also houses 4504691 SSR markers from all the breeds along with 1458 unique circRNAs, 37712 lncRNAs and 938 miRNAs. This comprehensive web-resource can be widely used by the buffalo researchers across the globe for use of markers in marker trait association, genetic diversity among the different breeds of buffalo, use of ncRNAs as regulatory molecules, post-transcriptional regulations, role in various diseases /stresses etc. These can also be used as biomarkers to address the adulteration and traceability. This resource can also be useful in buffalo improvement programs and disease/breed management.
Publications (2)
Whole Genome based web genomic resource for water buffalo (Bubalus bubalis).
Khan, Aamir Khan, Singh, Kalpana, Jaiswal, Sarika, Raza, Mustafa, Jasrotia, Rahul Singh, Kumar, Animesh, Gurjar, Anoop Kishor Singh, Kumari, Juli, Nayan, Varij, Iquebal, Mir Asif*, Angadi, U. B., Rai, Anil Rai, Datta, Tirtha Kumar, Kumar, Dinesh
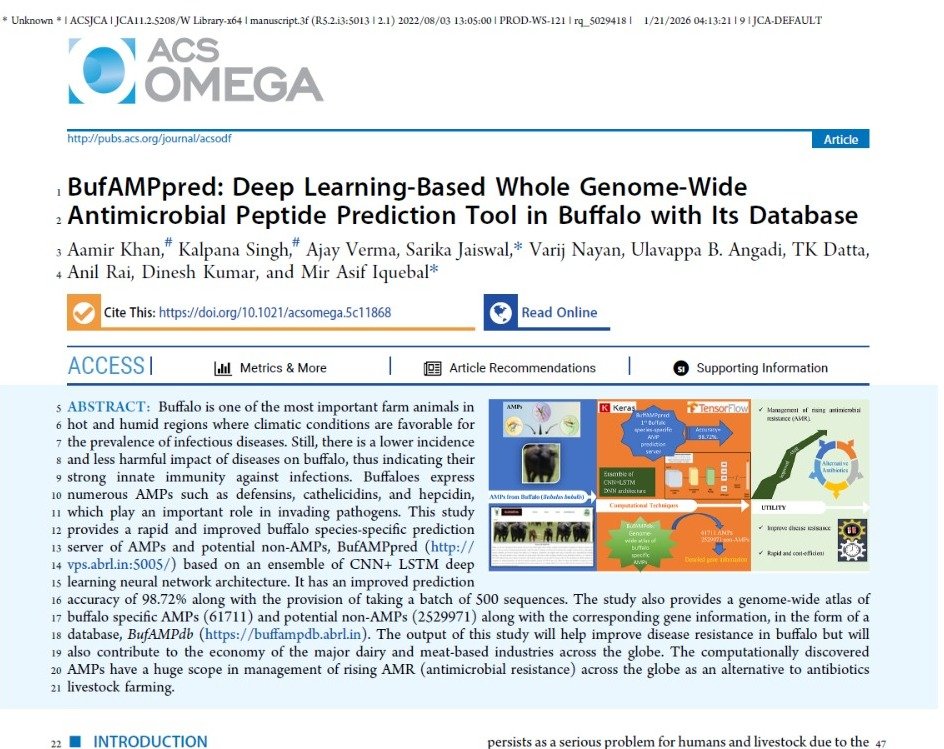

BufAMPpred: Deep learning-based whole genome-wide antimicrobial peptide prediction tool in buffalo with its database.
Khan, Aamir, Singh, Kalpana, Verma, Ajay, Jaiswal, Sarika, Nayan, Varij, Angadi, Ulavappa B., Datta, TK, Rai, Anil, Kumar, Dinesh, Iquebal, Mir Asif*